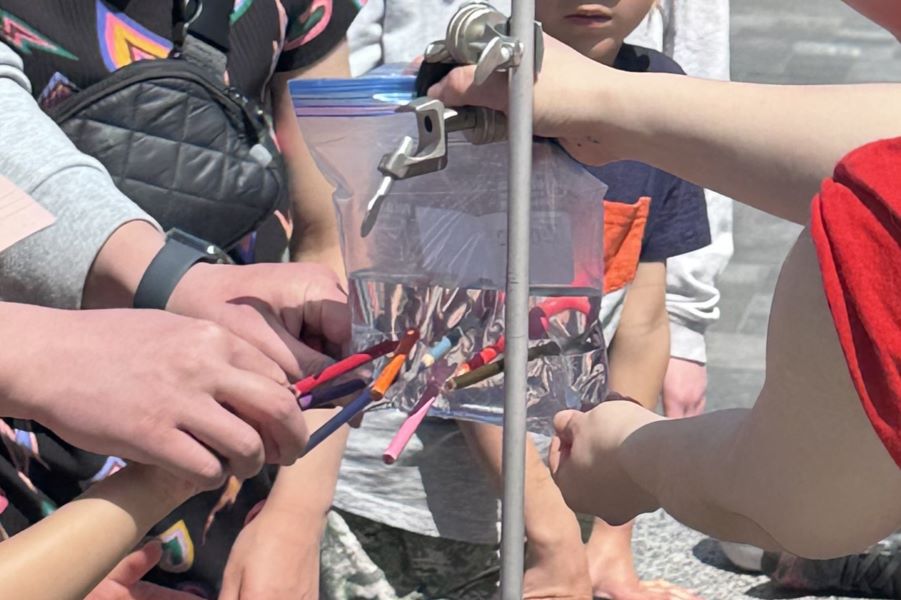Fresh excitement for the Science Rendezvous festival
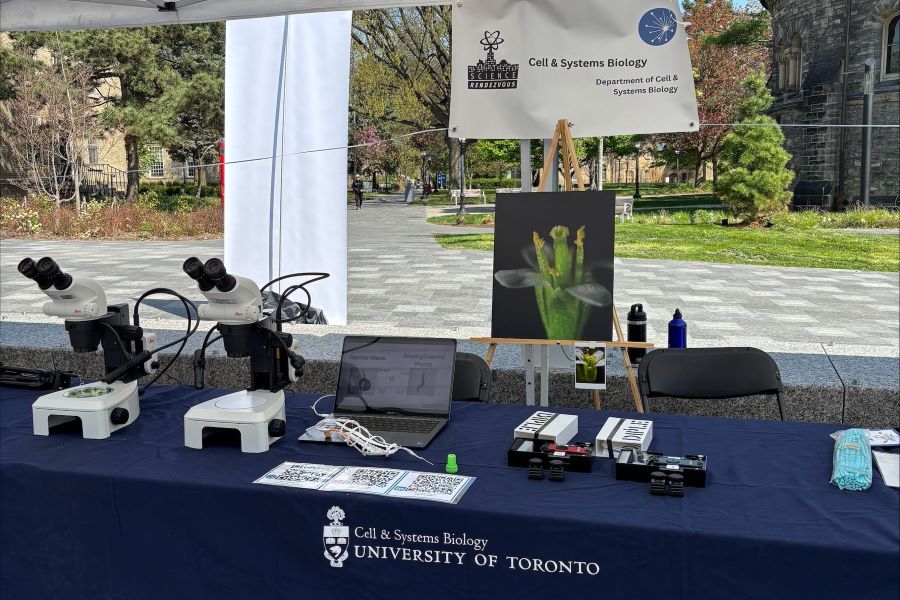
CSB shared our stories under shady trees as students, staff and faculty presented at Science Rendezvous 2025.
Science Rendezvous is an annual science festival across Canada, and we had many new faces from CSB to present our work on UofT's leafy Front Campus.
Squeezing in some neuroscience
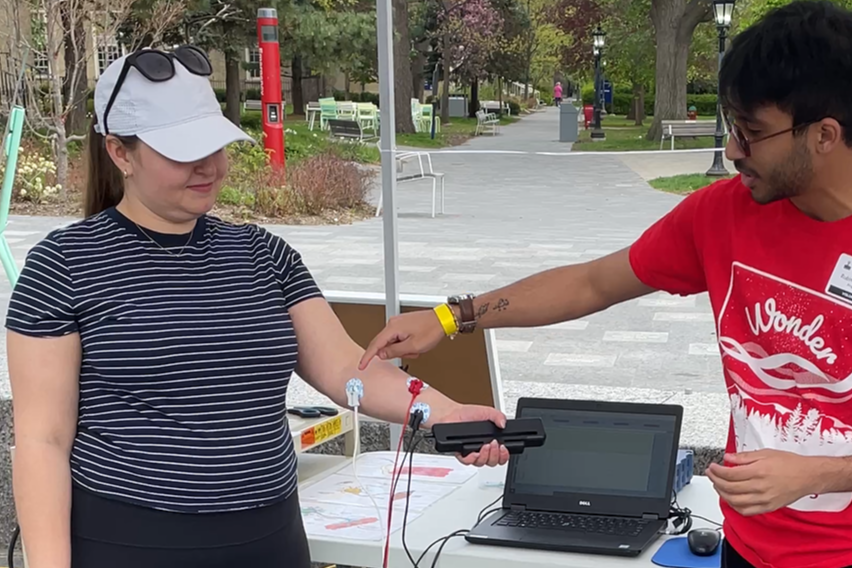
Stephanie Shishis works on behavioural neuroscience in the Kim lab. Aided by volunteer Rubin Khandekar and electrodes from the BIOPAC system, they measured visitors' grip strength while showing the electrical pulses that were activating the grip muscles. This led to discussions of how these impulses can go awry in diseases like Parkinson’s.
Linking shape and function at the molecular level
Matea Maurice demonstrated all the forms that proteins can take and the variations that can change function, drawing on her research in the Saltzman lab. Visitors made their own alpha helices, beta sheets and intrinsically disordered regions out of pipecleaners and took these model proteins home.
Victoria Zhang and Neil Macpherson demonstrated the different organelles inside our cells, each of which has a fascinating function. From the shipping warehouse of the Golgi to the powerhouse of the Mitochondria, Zhang and Macpherson showed our guests the complexity of life, even within a single cell. They helped visitors understand the shape these organelles make through forming their own organelles out of modelling clay.
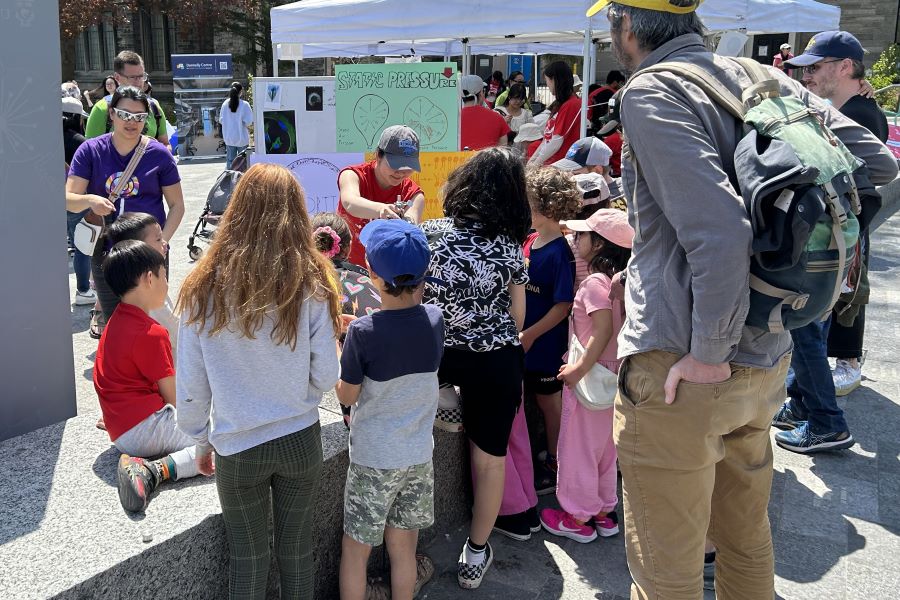
Karan Ishii from the Plotnikov lab cleverly demonstrated the physical forces that keep cells strong by showing how even a plastic bag that’s pierced all the way through won’t leak due to hydrostatic forces. She then explained to the large crowds gathered around her how she studies the forces exerted during cell migration and the cortical tension that keeps the cells intact.
It’s amazing what you can see with the right lenses
Plant scientists Professor Heather McFarlane and Yoshioka lab grad student Sofia Finley demonstrated the power of smartphone-compatible DIPLEscopes. With assistance from volunteer Abby Kuo, they revealed the columns, crescents and jigsaw pieces that cell walls make in different tissues of the plant; visitors took these beautiful patterns home on their phones!
Ernest Liang turned complex cellular patterns into a quiz. Guests were shown unlabelled microscopic images ranging from the folded leaves inside a seed to crawling cancer cells and were asked to guess which type of organisms they came from. The correct identification surprised many visitors, but some got it almost 100% right.
Our visitors were excited to look down the microscope and see tiny living organisms wriggling around on a plate. Ruby He and Linda Li of the Saltzman lab showed worms that move by rolling and going in spirals and contrasted these bioengineered worms with normal worms that move in a wiggly line. This activity was so popular, we had to cut off the line for visitors as our successful day came to an end.
Thank you to all our presenters, and to Lisa Matchett, Reta Aram, Tom Gludovacz and Alice DesRoches who helped with preparations!
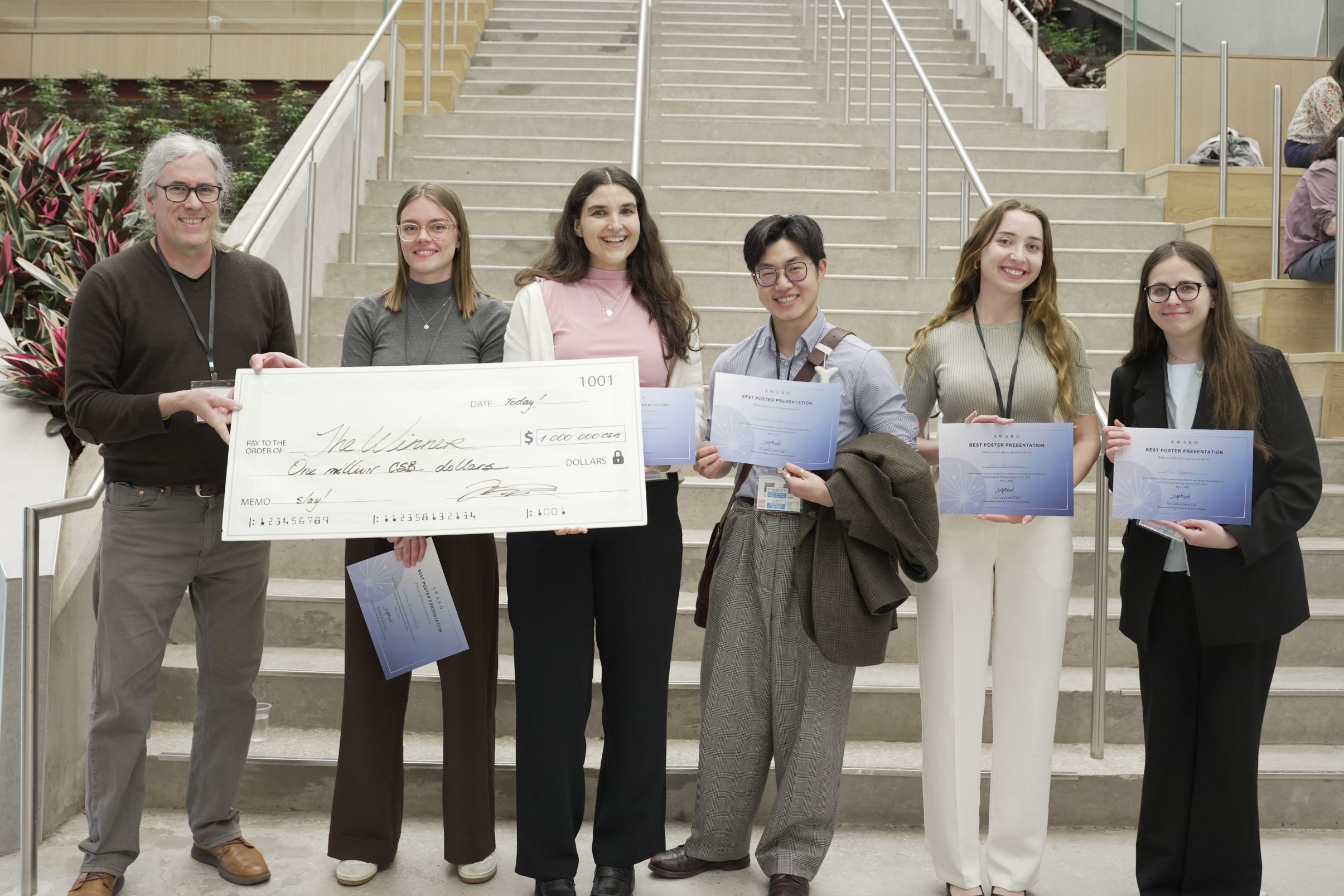
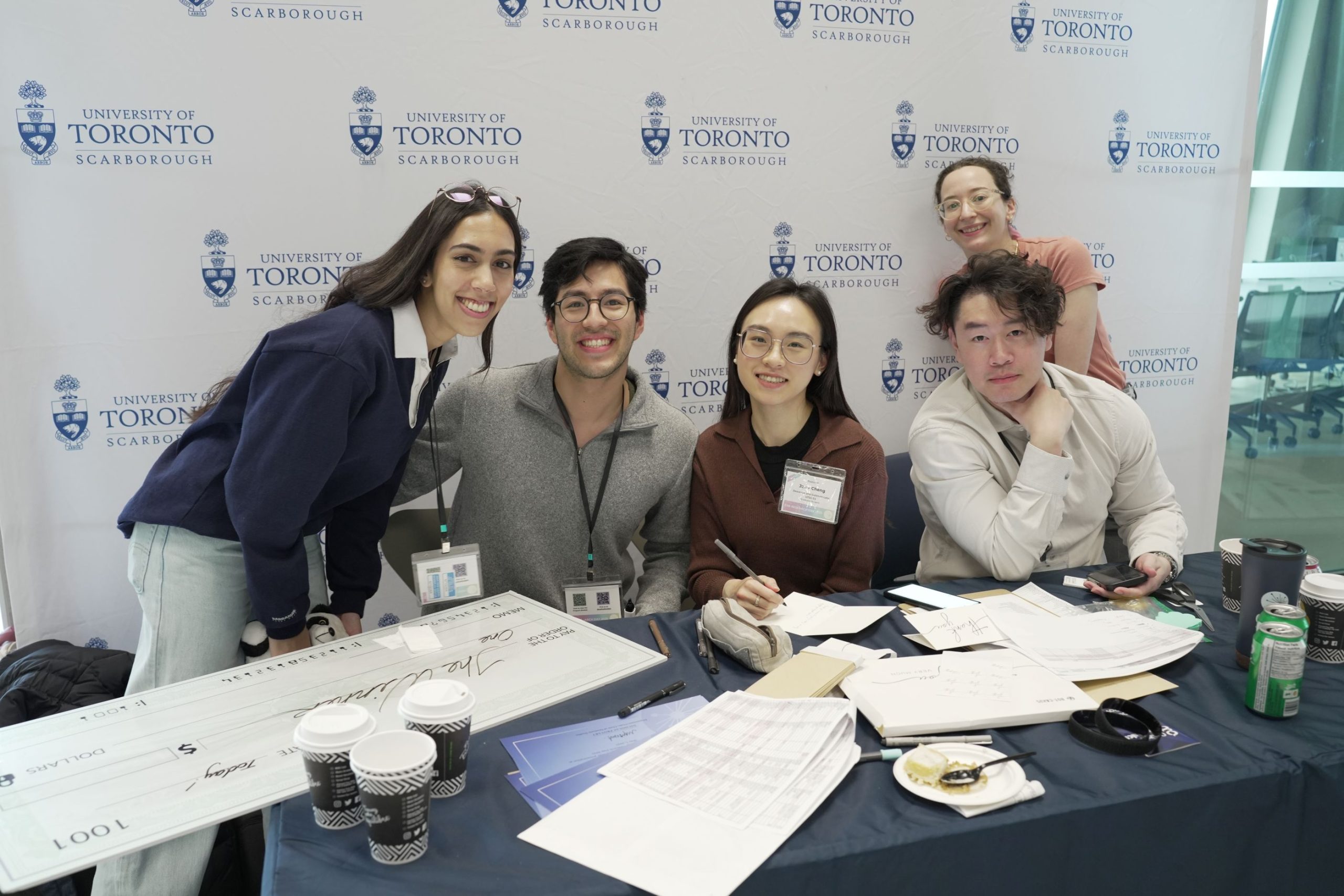
Innovation, Excitement and Awards at CSB Research Day 2025
CSB Research Day 2025 on May 1st at University of Toronto Scarborough gave us a chance to share fun times with our colleagues, gain important insights into their research and celebrate their accomplishments.
Students, staff and faculty were welcomed by Professors Rene Harrison and Adam Mott and introduced to our venue, the impressive Sam Ibrahim Building. Our keynote speaker Prof Costin Antonescu of TMU revealed fascinating details from his work on receptor signaling at the nanoscale.
From our experienced slate of speakers, two talks stood out as Best Oral Presentations: CSB Chair Nicholas Provart presented the awards to Brittany Dugan (Peever & Watts labs) for her talk on "REM sleep behaviour disorder in mouse models of Parkinson’s disease and multiple system atrophy" and Phoebe Bhagoutie (Saltzman lab) won her award for revealing "Heterochromatin readers CEC-3 and CEC-6 regulate the duration of transgenerational epigenetic inheritance in response to heat stress".
Some students presented their work in three minute Lightning Talks and the audience voted for Christine Nguyen (Nambara Lab) as giving the best talk for her presentation on "Characterizing the function of a novel abscisic acid transport inhibitor in Arabidopsis thaliana".
Most of our students presented their work in two poster sessions featuring animated expositions and excited queries. Judges assessed their research and presentations to award Outstanding Poster Presentations to
Norman Stewart (Ito lab)
Angela Sidsworth (Goring lab)
Mila Gorchkova (Anreiter lab)
Ernest Liang (Calarco Lab)
Irina Alymova (Peever lab)
Clare Breit-McNally (Desveaux & Guttman labs)
We were excited to welcome back alumni Luís Abatti, Laura Canales Sanchez, Sonhita Chakraborty, and Calvin Mok for our career and development panel. We are grateful for their participation and their thoughtful insights on life beyond graduate school.
The team organizing CSB Research Day and running registration did an amazing job. Our warmest thanks go to Raquel Singh, PJ Gamueda, Angela Sidsworth, Hasna Khan, Janis Cheng, Jiahao Chris Li (Chris also provided the photos in the gallery), Tamar Av-Shalom, Aiman Farheen, Rubesan Christian Joy Rajakumar, Denise Horsley and Professors Harrison and Mott.
We are grateful to our sponsors, New England Biolabs Canada, Ontario Genomics and TACT Genomics for their support.
Congratulations to our winners and thanks to everyone who presented their impactful work!
Prestigious teaching awards for two Professors from Cell & Systems Biology
We are proud that Professor Levy-Strumpf and Professor Yip in our Department have earned prestigious teaching awards in recognition of their excellence in education.
Professor Naomi Levy-Strumpf of the Human Biology program has earned the Cheryl Regehr Early Career Teaching Award from the University of Toronto. The Regehr Award recognizes faculty members who are effective teachers and who demonstrate an exceptional commitment to student learning, pedagogical engagement, and teaching innovation.
Human Biology Director Melody Neumann has high praise; “Professor Levy-Strump is an enthusiastic, energetic educator who demonstrates genuine and extraordinary care for her students’ learning and professional development, while inspiring others with her creative educational leadership.”
The award cites her dedication to teaching and commitment to furthering the student learning experience at U of T. Professor Neumann notes that “We’re very lucky to have her teaching in our programs and couldn’t be more pleased to see her work recognized with this award!”
Professor Kenneth Yip has earned the Outstanding Teaching Award – Early Career from the Faculty of Arts & Science. This award recognizes excellence in undergraduate and graduate education with a focus on classroom instruction, course design and curriculum development.
Nomination letters for Professor Yip “demonstrated widespread enthusiasm for your contributions and achievements as an outstanding teacher,” according to the award letter from Associate Dean Don Boyes. Boyes further expressed his “personal thanks for your ongoing commitment to teaching and to our students. Your contributions are greatly appreciated by this Faculty.”
Both Levy-Strumpf and Yip were previously chosen by students in Arts & Science for the Ranjini (Rini) Ghosh Excellence in Teaching Award, showing the broad appreciation their teaching generates from both students and Faculty.
Congratulations, Professor Yip and Professor Levy-Strumpf!
Gorgeous images win Nikon Small World Toronto competition
The winners of the Nikon Small World Toronto imaging competition have been announced!
We are proud that the top three winners as determined by imaging facility managers at SickKids Hospital were from Cell & Systems Biology! These images exploited the newest Nikon AXR NSPARC point-scanning confocal microscopy technology.
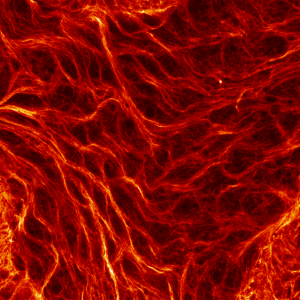
First place went to Damo Shi for “Immunofluorescence image of microtubules in the amnioserosa of fog-mutant Drosophila embryos”.
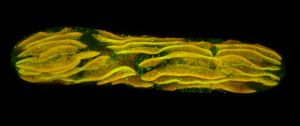
Second place went to Emily Deng for “From water to land: green algae transformed with an Arabidopsis protein required for chloroplast biogenesis”.
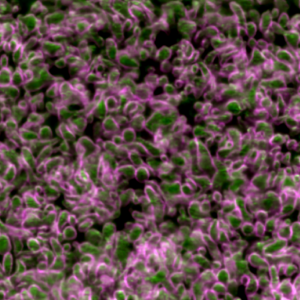
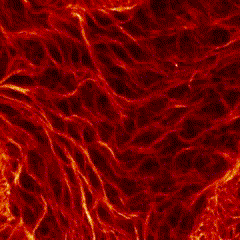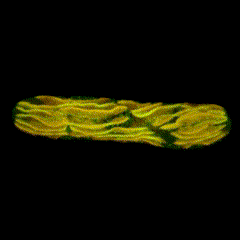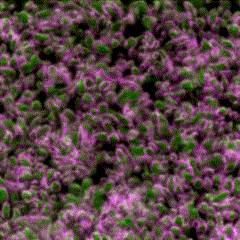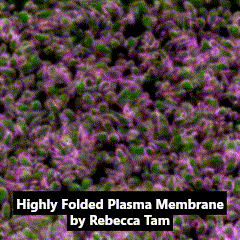
Third place went to Rebecca Tam for “Highly folded plasma membrane”.
You can see the full images in our gallery.
Shi’s research with Tirthankar Ray in the Harris lab showed that the amnioserosa of fruit fly embryos is confined by surrounding tissue to form a nematic, or crystal-like, structure.
Shi says that “My image for the competition submission was taken from an embryo with mutations of the gene fog, which perturbs confinement and causes reduced amnioserosa cell alignment.” Instead of aligning to outline crystal shapes, the red-stained microtubules in the fog mutant form stormy streaks.
Tam’s image shows a fluffy plasma membrane surface, contrasting with the usual image of the plasma membrane as a flat plane. This image comes from studies in early stage Drosophila embryos of how Arp2/3 proteins regulates cortical tension through patterning of myosin that is integrated with highly folded plasma membrane. In this image, the plasma membrane is coloured green and myosin is magenta.
Tam has shown how the plasma membrane is pulled by the centrosome to form a cap as fruit fly embryo cells divide, and this image shows the folding that occurs at a subsequent stage in the development of the Drosophila embryo. Tam recently won the Christine Hone-Buske award for her research.
Excellence in research shown at end-of-year poster session for undergraduate projects
Students in CSB497, CSB498, and CSB499 presented posters describing their 2024-2025 research projects at the end-of-year CSB Undergraduate Poster Session. Posters were assessed by judges from the Department and eight students earned the F Michael Barrett Award for their excellent presentations. The winners were presented with their awards by Undergraduate Chair Prof Dinesh Christendat.
Students studying model animal systems studied the role of growth factors in stem cells, the effect of radiotherapy on tissue, and made discoveries in gene regulation at the level of both DNA and proteins.
Cindy Yu Fei Lei (Gilbert Lab) won for "In Vitro Characterization of FGF2 as a Muscle Stem Cell Niche Occupancy Modulator"
Caitlin Hui Xarn Tan (Yip lab) presented her results on "Exploring the Role of Tet2 Chip Mutation in the Development of Radiation-Induced Fibrosis"
Yaqing Zhao (Mitchell lab) revealed her insights into "Using Synthetic Transcription Factor Libraries to Decipher the Enhancer Code"
Grace Huang-Zhan (Saltzman lab) was recognized for her "Characterization of the Developmental Delay in C. elegans Polycomb Repressive Complex 1 Mutants"
Undergraduate plant scientists revealed their insights into plant immune responses at the transcriptional and organismal level, as well as probing biochemical regulation of stress responses.
Wan Ni Chow (Yoshioka lab) determined the "Temperature Dependence of Direct Antifungal Effects in ISR-Inducing Plant Growth-Promoting Rhizobacteria"
Reid William James (Yoshioka lab) detailed their work "Investigating the Transcriptional Network within the Cyclic Nucleotide-Gated Channels (CNGCs) in Arabidopsis thaliana"
Vy-nhan Chanh Nguyen (Nambara lab) was recognized for "Unravelling the Regulatory Mechanism of ABA Catabolism Under High Humidity in Arabidopsis Thaliana"
Nicholas Garcia (Guttman lab) presented his work on unique transcriptomic responses evoked by PtoDC3000 Type 3 Secretion Effectors in Arabidopsis thaliana
Congratulations to these winners, and to all poster presenters for their hard work! Thank you to Melissa Casco and Genna Zunde for their hard work organizing the poster session.
Rising star in plant sciences Heather McFarlane awarded CSMB New Investigator Award
CSB Professor Heather McFarlane has earned the prestigious New Investigator Award from the Canadian Society for Molecular Biosciences (CSMB).
The CSMB grants this award to rising stars in cell and molecular biology and this award recognizes McFarlane's outstanding research on plant cell wall synthesis. Plant cell walls provide structure and flexibility to plants that allow them to form thin, flexible structures like corn stalks or trees.
Stress on plant cell walls, including stress from climate change, can reduce yields in important products like lumber, cotton and fuel crops. Previous attempts to modify plant cell walls for improved materials or biofuels have resulted in substantially reduced crop yields, showing that we don't fully understand how this economically important process occurs.
The goal of McFarlane’s award-winning research is therefore to study how plants sense and respond to cell wall changes to permit cell wall engineering that advances sustainability and yields innovative products. McFarlane and her trainees have already revealed important aspects of cell wall synthesis.
They have identified dozens of new molecular components of cell wall signaling and provided mechanistic insights into why some cell wall modifications are tolerated by plants while other changes can cause dramatic growth and developmental phenotypes. We look forward to further revolutionary work from this team.
McFarlane's excellence is reflected by her post as CRC Chair in Plant Cell Biology, by her Ontario Early Researcher Award and by her earning the Dorothy Shoichet Women Faculty in Science Award of Excellence.
Congratulations, Heather!
Congratulations to Professor Woodin on her appointment as President of University of Toronto
Professor Melanie Woodin, an internationally recognized neuroscientist at CSB, has been named the University of Toronto’s 17th president!
An accomplished leader, Woodin is current Dean of Arts & Science at U of T, president of the Canadian Association for Neuroscience and serves on the board of directors at the Vector Institute.
Woodin's Lab at CSB is focused on discovering the cellular mechanisms underlying inhibitory GABAergic synaptic plasticity in the healthy and diseased brain.
They employ a multi-disciplinary approach combining electrophysiology, biochemistry, fluorescence imaging, and behaviour to understand how synapses are built, the mechanisms that underlie their plasticity, and their role in neuronal circuits.
Woodin's studies unravel mechanisms that lead to neurological disorders and diseases, including autism spectrum disorder, Huntington’s disease and ALS.
“I am profoundly honoured to accept this appointment to lead an institution that I care about deeply,” says Woodin. “U of T is widely recognized as one of the world’s best universities and a highly trusted source for ideas, research, innovation and talent.”
Congratulations, Melanie!
Laptop or Lab? Research opportunity in Chang lab leads BCB student to embrace both
A Research Opportunity in the Chang lab led Alena Qin, a student in the Bioinformatics and Computational Biology program, to an appreciation for hands-on experimentation.
Students in Arts & Sciences can engage in research opportunities through the Research Opportunities Program. Qin was accepted into the lab of Professor Belinda Chang, whose lab works on recreating the evolution of visual proteins in the laboratory, using dry-lab computational methods, but also wet-lab mutational studies.
“The ROP has definitely helped me narrow the scope of the type of research I'm interested in,” say Qin. “I’m really enjoying the wet lab experience and I'm thinking of pursuing a career that’s more in line with that.”
You can read more about Qin's experience in "A&S Research Opportunities Program provides second-year undergrad Alena Qin with hands-on lab experience.
ROP courses are posted on CLNx in February when the applications start. You must apply on CLNx within the program application timeframe, which is from mid-February to mid-March every year, to be considered.
For other positions in Cell & Systems Biology, students can review the list of Faculty for labs researching their topics of interest and contact the professors. Not all professors have available positions in their labs, so it's good to contact several professors.
We are glad to offer these research positions to help students learn where their future can take them.
Excellence in Plant Science earns Professor Heather McFarlane the Dorothy Shoichet Award
 Congratulations to CSB Professor Heather McFarlane on earning the Dorothy Shoichet Women Faculty in Science Award of Excellence. McFarlane is receiving the award to support her research into how plants sense and respond to cell wall changes, work that has important applications in the production of renewable bioproducts including paper, cotton, wood, compostable bioplastics and next-generation renewable biofuels.
Congratulations to CSB Professor Heather McFarlane on earning the Dorothy Shoichet Women Faculty in Science Award of Excellence. McFarlane is receiving the award to support her research into how plants sense and respond to cell wall changes, work that has important applications in the production of renewable bioproducts including paper, cotton, wood, compostable bioplastics and next-generation renewable biofuels.
"Heather is a stellar researcher who produces world-class research through ingenuity and thoughtful guidance for trainees in the lab." says Professor Jennifer Mitchell, CSB's Associate Chair, Research. "This award will accelerate the pace of her discoveries by allowing her to devote more time to her research with long term impacts in sustainable agriculture, food security, and next-generation biofuel development."
The Dorothy Shoichet Women Faculty Science Award of Excellence was established in 2016 by University Professor Molly Shoichet in honour of her late mother. It provides release from teaching for early-career, female researchers in any of the physical or life sciences, computer sciences or mathematics within the faculty.
“I'm deeply honoured to receive the Dorothy Shoichet Award,” says McFarlane. “Time is a valuable resource in the lab and this award provides precious time and the opportunity to focus on research and research training. The award will accelerate my lab's research as we pursue several exciting directions in our efforts to understand plant cell-wall signaling."
Congratulations, Prof McFarlane!
Supporting the Talented Women Scientists in CSB on IDWGS
We are proud to recognize the accomplishments of our women colleagues on 2025's International Day of Women & Girls in Science and to offer our support all year long.
This year saw many accomplishments from women scientists at CSB! Follow the links below for more details on some of these achievements:
Professor (and former Chair) Daphne Goring earned a Gold Medal awarded by the Canadian Society of Plant Biologists for a career of outstanding contributions to plant science.
CSB student (now Dr) Sonia Evans resolved the long-standing "pyruvate paradox" in a publication at Nature Plants by discovering the pathway whereby pyruvate is generated from rubisco in the chloroplast. Her outstanding skills earned her a Provost Post-Doctoral Award from UofT.
Professor Heather McFarlane showed how building the economically crucial plant cell wall is dependent on correct labeling of its components in a publication at Development Cell. Her excellence was recognized in the renewal of her Canada Research Chair in Plant Cell Biology.
Teaching Assistants are a vital part of our courses and this year, four women received commendations from life sciences undergraduates for their outstanding skills. Congratulations to Ruby He, Mary-Elizabeth Raymond, Andrea-Aditi Taylor and Kathryn McTavish on your TA Teaching Excellence Awards!
CSB is committed to leadership and innovation in research and teaching. We incorporate equity, diversity, inclusion, and integrity in our operations. We value respect, professionalism, and collaboration in our community. None of this is possible without acknowledgement and support for women and girls in science.