Research Day 2026 reveals astonishing advances from CSB across life sciences
CSB Research Day 2026 was held on May 1st at UTM's Kaneff Centre. This event showcased impressive innovation across the kingdoms of life.
Our keynote speaker, Professor Sheena Josselyn of SickKids Hospital presented her thought-provoking research on how memories are formed and erased in a Taylor Swift-themed talk.
We are grateful to Ryan Ruan (BSF) and Dr Burton Lim (ROM) who provided important job perspectives in a lunchtime career panel.
Through talks in the KN137 lecture theatre and posters in the Rotunda, CSB graduate students clearly and concisely presented their discoveries. Some presentations were chosen by judges as outstanding efforts and earned awards at the end of the day, presented by N. Ross Stewart and Bauhua Liu of the Organizing Committee.
Best Spotlight Talk Awards
Claire Fernandes (Guzzo Lab) for "Emerging Cytometry Techniques To Purify And Study Virus Subpopulations"
Milena Russo (Liu Lab) for "Circuit Mechanisms Underlying Global Motion Processing in the Brainstem"
Best Lightning Talk Award
Anthony Kadamani (Peever Lab)
Best Poster Awards
Matthew Danesh (Peever Lab)
Congrong (Ruby) He (Calarco Lab / Zhen Lab)
Serene Moussaoui (Terebiznik Lab)
Ryan Ugovsek (Lin Lab)
Mackenzie Wilson (Phillips Lab)
Congratulations to these talented presenters! Thank you to the Organizing Committee N. Ross Stewart, Bryan Guo, Gary Chatha, Boahua Liu, Adam Mott, Shelly Lumba, Ben Eldridge and Denise Horsley. Photographs by Angela Sidsworth.
Epigenetic markers of neuronal gene regulation revealed to earn Hone-Buske award
Kailynn MacGillivray has earned the Dr Christine Hone-Buske Award for her outstanding publication “Widespread association of Polycomb complex–deposited histone H2A monoubiquitylation with enhancers and neuronal gene regulation”. This research in Science Advances reveals the role of protein modifications in the developing nervous system of the worm C. elegans.
MacGillivray’s work in the Saltzman lab focused on ‘epigenetic’ changes promoted by the protein complexes PRC1 and PRC2. These changes act on histone proteins bound to DNA, serving to turn genes off by packaging them tightly or turn them on by opening up the DNA surrounding the gene.
MacGillivray focused on ubiquitin and methyl molecules attached to histones. “We observed what we thought was a pretty unique phenotype in the worms if you look at PRC1 mutants for C. elegans versus PRC2 mutants.” MacGillivray explains.
“We found that genome-wide these histone modifications weren't working together in the same way as in mammals or fruit flies. Overall methyl patterns were unexpectedly maintained when ubiquitin was depleted.”
Given this unusual observation, MacGillivray used genome analysis to reveal that genes misregulated in ubiquitin-deficient mutants are enriched for nervous system functions including neuronal differentiation and axon guidance.
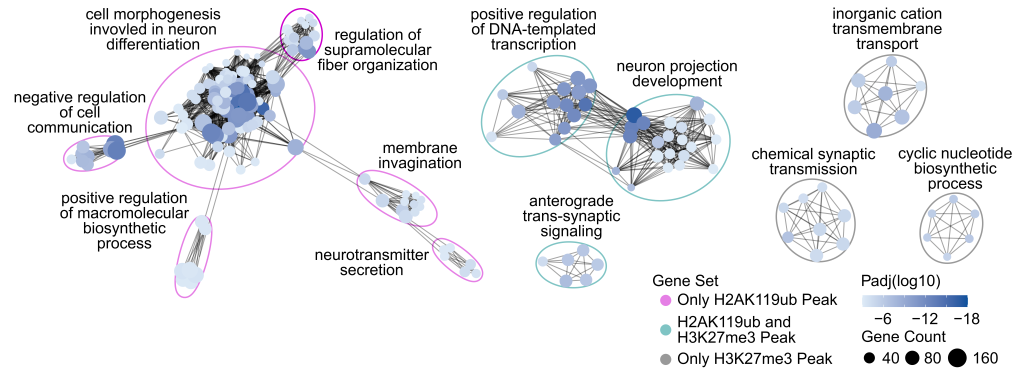
An exciting insight from this analysis was that epigenetic changes in PRC1 mutants were enriched at ‘enhancer’ regions: sequences outside of genes that regulate where and when a gene is expressed, even at a distance.
MacGillivray was puzzled: “We had this enrichment at enhancer regions and at neuronal genes, but we didn't have an obvious link in terms of a gene regulatory standpoint how these two are related in the developing worm. Then we were looking through recent papers and saw a set of enhancers that become nicely accessible or potentially active within neuronal tissue.”
“I found that ubiquitination was most highly enriched at these enhancers compared to enhancers that are accessible in other tissues,” MacGillivray explains. “That was definitely the ‘oh, this makes sense!’ moment that tied this paper together.”
There was excitement in the Saltzman lab as MacGillivray showed the results to her colleagues and mentor. Their animated discussions postulated that epigenetic changes drive temporal control as the nervous system develops and this may be what is constraining activity.
The unique aspects of worm neuronal development can provide insights into other organisms. Some neurodevelopmental dysfunctions in humans are linked the PRC1 complex, and the Saltzman lab is collaborating with researchers in Miami to see if worms provide a model for understanding these changes in humans.
“It’s been great to follow all these different pieces of evidence,” says MacGillivray. “I’ve always been curious, and making these discoveries to me is the whole point of science. I’m glad to have this enthusiasm rewarded with the Dr Christine Hone-Buske Award.”
Congratulations, Kailynn!
Intensive Laboratory Experience prepares Life Sciences students for a career in research
The Human Biology Lab Bootcamp is an intensive two-week wet and dry lab training program offered to students finishing their third or fourth year of studies in life sciences. It provides hands-on research experience, technical skill development, and guidance for students considering careers in biomedical research or related fields.
The HMB Program will host the 2026 Lab Bootcamp on May 4–15. This no-cost, ungraded program is designed for students with fewer opportunities to gain hands-on biology lab experience. The Bootcamp was conceived and designed in 2017 by University President Dr Melanie Woodin (then Director of HMB) by Dr Colleen Dockstader.
Unlike weekly lab classes, each day’s experiments build on the previous day’s work—like chapters of a story—so students follow multi-day workflows. Topics include DNA purification, restriction digestion, cloning, gel extraction, ligation, colony PCR, tissue culture and transfection, immunofluorescence microscopy, and more.
Groups of eight students are supported by one facilitator. Those accepted for the program are always looking to maximize their skills and knowledge without the stress of regular tests.
The program gives special recognition to Drs Colleen Dockstader, Jasty Singh, and Alan Wong for creating the lab manuals and designing this Bootcamp in 2017.
As this program progressed, professors and postdocs helped sustain the program, including Drs Jessica Pressey, Naijin Li, and Samuel Delage, as well as many facilitators who taught students and guided them through the course each day.
We look forward to welcoming this year's cohort of dedicated students!
Prof Haley Zubyk Earns ASSU Award for Excellence in Teaching
Congratulations to Professor Haley Zubyk, who was nominated by Arts & Science students to receive the Ranjini (Rini) Ghosh Award for her Excellence in Teaching!
The award was earned by Zubyk’s ability to stimulate and challenges students’ intellectual capacity, skill at communicating the course material to students, mastery of the subject area and for being highly accessible to students
"Receiving this student-nominated award is incredibly meaningful to me because it reflects that my students feel supported throughout their undergraduate journey and that my passion for teaching is visible to them,” Zubyk says. “Teaching and learning are deeply rewarding, and my students are without question the best part of this work.”
Zubyk's students experience her passionate pedagogy studying the Challenges of Antimicrobial Resistance in Human Biology and attending her course on AIDS: A Global Perspective in the Fall term and in Summer sessions. In the Winter term, Zubyk presents an Introduction to Human Biology and shares insights into The Human Microbiome in Health & Disease.
Congratulations, Professor Zubyk!
A year of deep research leads to a culminating event for fourth year students
Fourth year students in CSB can join our research laboratories to conduct projects in their favourite topic as part of CSB497, 498 or 499 courses. Their final task after generating data and analyzing results is to present their conclusions at our annual Undergraduate Poster Session.
This year, the poster session was held on March 27th and featured dozens of posters showing students' discoveries in cell biology, cancer biology, developmental biology, neurobiology and plant biology.
The posters were judged by departmental staff and faculty. They assessed the content of the posters and the presenters' ability to describe their projects and answer questions.
The F Michael Barrett Award was presented to students who excelled at their presentation by Undergraduate Chair Professor Keiko Yoshioka.
Faten Abla (Bruce lab) showed their expertise in "Investigating the Role of Eph/Ephrin Signaling in Tissue State Transitions During Zebrafish Mesoderm Internalization"
Maia Edney (Yip Lab) pursued "Investigation of Novel Compound for Suppression of Extracellular Matrix Proteins"
Alex Huang (Plotnikov Lab) demonstrated how "ER Ca2+ Regulates ER: PM Contacts to Facilitate Linear Migration in Elevated ECF Viscosity"
Tiantian Lei (Lin Lab) studied the "Effects of GCaMP6 on Effects of GCaMP6 on the Survival and Behaviour of Zebrafish Larvae"
Aliya Mohd Zamri (McFarlane Lab) took on the task of "Confirming putative cell wall signaling mutants from a forward genetic screen"
Shannon O’Reilly (Calarco Lab) demonstrated a "High-Throughput Behavioural Monitoring Platform to Identify RNA-Binding Proteins Regulating Locomotion in C. elegans"
Zoe Smith (Yip Lab) presented results from "Investigating the Role of FOXO1 in Radiation-Induced Fibrosis"
Tea Reed Watson (Lumba Lab) showed how "ShHTL7 mutants demonstrate altered germination and temperature sensitivity compared to wild-type in an Arabidopsis model"
Fiona Yi Yang Xu (Bruce Lab) explored "Frmd4 Proteins at Cell Junctions: Potential Cytohesin Adaptors during Zebrafish Epiboly"
Vicky Ziyi Wei performed her work in the Tepass lab, which studies the cellular and molecular mechanisms that control the polarity of epithelial cells and the cell adhesion between epithelial cells that binds cells into tissues
Congratulations to these impressive students on their accomplishments and good luck as you move on to graduation!
Prof Keiko Yoshioka earns Manery Fisher Award from CSMB for research and leadership
Professor Keiko Yoshioka has earned the Jeanne Manery Fisher Award from the Canadian Society for Molecular Biosciences in recognition of her excellence in research, teaching, mentorship and inclusiveness.
Yoshioka's strengths are shown in two fields of research; plant immunity signaling and crop protection using natural stimulants. She is a pioneer in identifying a group of membrane calcium channels, known as cyclic nucleotide-gated ion channels (CNGCs), as key regulators of plant calcium signals during stress responses. Her research on using non-pathogenic microorganisms to sustainably protect crop plants, and thereby food production, is transformative.
The Plant Genomics and Biotechnology focus at CSB was conceived and co-led by Yoshioka as part of promoting the importance of plant research to UofT undergraduate students. Yoshioka also acts as Undergraduate Associate Chair of CSB.
Over 200 undergraduate and graduate students have worked in the Yoshioka lab, and the quality of her mentorship has led past students to return for professional advice, and prompted some to become professors. Recruitment to her lab is guided by EDI principles.
Yoshioka's support for inclusiveness led her to co-ordinate conferences in Toronto on gender equity and diversity in science, as well as supporting scientists in Africa combating plant diseases through training and research.
This remarkable career makes Yoshioka eminently deserving of the Jeanne Manery Fisher Award in honour of a ground-breaking biochemist and teacher who promoted equity throughout her career.
Congratulations, Keiko!
Many pathways to an exciting career revealed at the CSB/HMB backpack to Briefcase event
Thank you to all the alumni who answered the call to share their career paths and job advice at our "backpack 2 Briefcase" event with HMB and CSB undergraduates!
What are the career options for CSB and HMB students?
After introductory remarks from Professor Eba Kurz, students travelled between tables at the Wiliam Doo auditorium to ask questions of different alumni.
“I got to meet a lot of different people who are intersectional in terms of their career in biology and something else,” noted one student “So it's just been fun to explore all the different career pathways I could potentially take.”
Our guests were at different career stages from entry level through research and service professionals to senior managers. Positions in academia, pharmaceuticals, finance, and public service were all represented. They had roles as research scientists, consultants, lawyers, sales specialists or clinic directors. This diversity gave each alumni unique perspectives on the paths open to our students.
One of the students learned that it was important to “Explore your options because if I was in first or second year and this event came up into my e-mail I wouldn't have looked out for it. But with my experiences into my third year, I think the most important thing I can take from undergrad is go for the opportunity and not just wait for it to fall in your lap.”
Khevna Dave studied Global Health and worked in vulnerable communities in South America, West Africa, South Asia and Middle East. She is now back in Toronto as Director of Mission Hub, training people on how to do cross-cultural work well.
“A lot of volunteering opportunities as a student here actually played a role in developing some of my skills in my current work,” Khenva asserts. “I volunteered with extra-curricular clubs that built my networks and connections with people. And you can work for UofT in different roles as a student. I did a lot of communications at Student Life that naturally lent itself as a helpful tool for my nonprofit work.”
Many pathways to an exciting career
One student learned that “Our pathway is not just set into stone. The first speaker that I talked to actually was telling us about how her undergrad differed a lot from her postgrad with her masters and where she's working now. And that really reignited a lot of my previous passions that I haven't considered.”
Trieu Nguyen was one alumni who took an alternative path. “I started my journey in Human Biology, taking all the different science courses for the first three years. And then I kind of did a little pivot in my undergraduate program. I took a little bit of statistics and political science and ended up falling in love with computers. So you can start in one area, but you can always try something else and venture into different areas. I now run a team of developers and business analysts as clinical research coordinator at Bank of New York.”
Reflecting on an informative experience
This event proceeded smoothly thanks to the work of faculty and staff from Faculty of Arts & Science: Genna Zunde, Mila Thirasack, Sarah Cook, Prof Eba Kurz and Prof Keiko Yoshioka.
At the end of backpack to Briefcase, students were positive about the experience “I came here looking into what I want to do for grad school or med school. Seeing how people have progressed through their career is kind of what I'm looking for. I found out about a bunch of different paths through and after life science. So I definitely got what I came for.”
“I learned a lot about how to talk to your professors and make sure that you can get a good reference and then also just how to make your application to different grad schools stand out more,” said one student guest. “But I think this is my first event where I got to speak with a lot of different alumni. I'm really glad that I came to this event today.”
The alumni were impressed by what they saw as well. Max Shcherbina studied Cell & Molecular Biology and is now a Research Scientist at University of Toronto “What struck me most about the students who came is how eager and how wonderful their achievements are so far and how they want to keep that going. And I was happy to facilitate and help as much as I could in that regard.”
Thank you to Dr Meggie Cao (Cell & Systems Biology), Max Shcherbina (Cell & Systems Biology), Dr Simon Lui (Nikon), Marissa Lee (Massage Addict), Limin Wang (OLG), Sabrina Hyde (Ontario Ministry of Children, Community, & Social Services), Ahash Jeevakanthan (Public Health Canada), Trieu Nguyen (Bank of New York), Katelyn Macfaden (Borden Ladner Gervais Llp), Rabia Ahmad (Supply Ontario), Khevna Dave (Mission Hub) as well as Muna Abdulla (Trillium Health Partners), Tonny Huang (GSK), Dr Arnold Ruste (PrEP Clinic) and Thenuka Thanabalasingam (Sanofi)!
Secrets of gene regulation during development probed with new tools by Professor Colette L. Picard
Professor Colette L. Picard is bringing her innovative skills in single cell genomics to the Department of Cell & Systems Biology (CSB) as a new professor at University of Toronto. She was drawn to a city packed with life science innovators, and to CSB in particular for our deep expertise in bioinformatics and genomics.
Precision resolution and innovative computational techniques in gene regulation
Picard is a pioneer in single cell analysis of developing tissue. She was one of the first scientists to use new single cell sequencing techniques to probe gene expression in complex tissues in plants, focusing first on the developing seed. This study found that an unusual gene expression pattern called imprinting is elevated in a small region that forms the interface between the seed and the mother plant, highlighting the level of detail that these new single cell sequencing approaches can capture.
Single cell sequencing uses sophisticated high-throughput, low-input techniques to prepare sequencing libraries of thousands of individual nuclei simultaneously. Analyzing these complex datasets poses a number of challenges, and Picard relies on her computational and statistical background to tailor each approach to her biological questions.
Picard is intrigued by how gene expression is regulated, particularly during development. Her studies have converged on epigenetic alterations to DNA, particularly the addition of methyl groups to DNA. Epigenetic modifications alter the expression of the affected regions of DNA. Since all the cells of a developing organism are genetically identical, epigenetic modifications are one way that this single DNA ‘blueprint’ can give rise to a large variety of cell types and behaviors within a tissue or complex organism.
An important model system for the very start of the developing organism
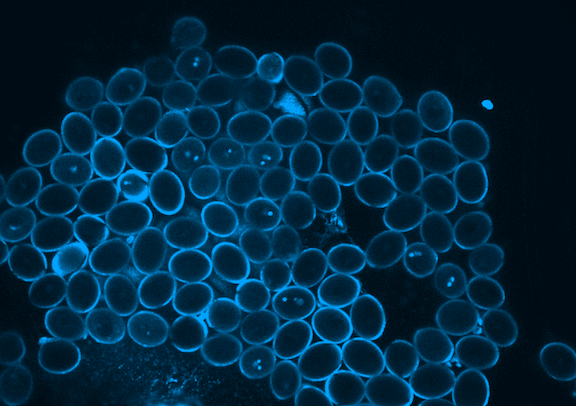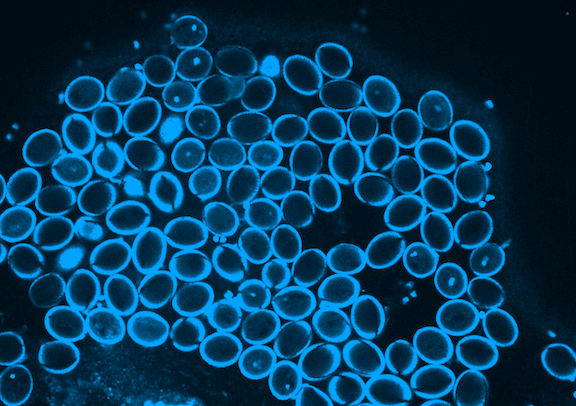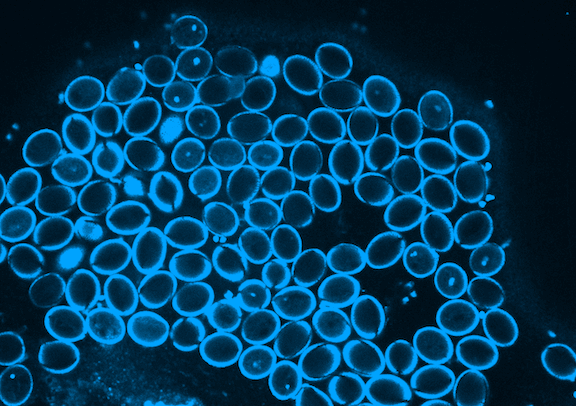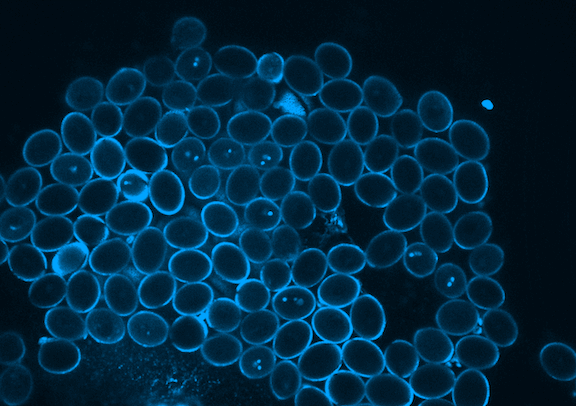
Picard is currently focused on the development of the germline, which in plants occurs when the pollen is formed. Pollen is made of three genetically identical cells, each with distinct behavior and gene expression patterns. Picard is interested in understanding the sequence of epigenetic changes that create the differences between these three cells. In fact, pollen is an excellent model system for developmental biology: it’s simple, abundant, and has only one copy of every gene instead of the normal two.
But it’s not just a great model system; pollen is also important to study. Yields of major crops like corn and wheat depend on pollination. Across the entire lifecycle of the plant, pollen development is particularly sensitive to environmental stress. A heat wave just as pollen grains are forming greatly reduces the formation of viable pollen. Picard will probe the effects of extreme heat on gene expression during pollen development to understand and propose solutions to crop yields reduced by extreme heat.
Growing a lab in Toronto
Professor Picard is currently accepting graduate student applications from those interested in studying gene regulation and epigenetics in germline development. Her lab will consist of both a ‘wet-lab’ equipped to grow plants and run experiments on pollen, as well as a ‘dry-lab’ focusing on the analysis of high-dimensional datasets like single cell sequencing data in a variety of plant and animal systems.
Picard is grateful to her postdoc advisor Steve Jacobsen at UCLA and her PhD advisor Mary Gehring at MIT for their support in her successful application, and looks forward to joining the vibrant scientific community at CSB. She is enjoying getting to know Toronto and reassures us that despite moving here from Los Angeles, she is originally from Michigan and Montréal and is comfortable with the blizzards obscuring our streets.
We are excited to see the discoveries that will come from her research! Welcome, Colette!
Technical innovations in the Plotnikov lab reveal machinery behind cell migration in unprecedented detail
An international team led by a UofT lab has revealed a novel regulatory system behind cell migration and shown how the different parts work together.
This publication in the journal Current Biology adds to our understanding of how healthy bodies are maintained, as cell migration is crucial to the developing body and to immune cells traveling to the site of injury.
How do cells push out into their environment?
Ernest Iu, a graduate student in Cell & Systems Biology, decided to probe how calcium controls the structure and dynamics of the actin filaments at the leading edge of migrating cells.
Actin filaments are rods that grow within the cell to push out against the cell membrane, leading to flaps called ‘lamellipodia’ that promote movement. Scientists for years have noticed the importance of calcium ions in the regulation of lamellipodia dynamics, but how exactly this regulation occurs has remained unclear.

In a technical tour-de-force, Iu and his colleagues in the Plotnikov lab exposed novel details of what’s happening under the hood of calcium-dependent cell migration. This work is published as “A TRPV4-dependent calcium signaling axis regulates lamellipodial actin architecture to promote cell migration”.
Iu’s supervisor, Professor Sergey Plotnikov, is enthusiastic about this work: “Previous investigations into the biological roles of calcium signaling pathways have been largely limited to observing cellular responses. We have revealed the machinery behind a long-sought signaling mechanism that regulates the actin cytoskeleton in cell migration.”
Technical innovations demonstrate cellular machinery in unprecedented detail
Cell migration is usually observed on cells growing on glass, but the ragged edges of the cell make quantitative analysis difficult. Iu’s technique circumvented this shortcoming by observing round cells as they settled onto the bottom of the dish and spread out in a circle.
He showed slower spreading in cells without the TPRV4 channel compared to control cells, demonstrating that this calcium channel was important for lamellipodia. Iu then proceeded to look inside the cell to reveal an intricate mechanism of interacting proteins dependent on TRPV4.
Initial results showed that inhibiting TPRV4 inhibited the RhoA protein. “I thought this was a dead end,” states Plotnikov. “How do you study the functional connections of two seemingly unrelated proteins? But Ernest was too stubborn to stop.”
Iu’s perseverance first identified a molecular switch that controls other proteins, known as a kinase CaMKII. CaMKII that becomes active when the TRPV4 channel is activated.
Next, Iu innovated a technically challenging procedure to isolate proteins from the migrating region of the cell. Iu and colleague Alexander Bogatch placed the cells on a fine mesh and provoked the cells to grow lamellipodia through the mesh.
They then collected these microscopic protrusions to isolate the proteins at the tip for mass spectrometry analysis. The success of this advanced cell fractionation and mass spectrometry experiment was the result of a global collaborative effort with the Humphries lab in Manchester, made possible by generous support from the University of Manchester-University of Toronto Joint Research Fund (now the MMT Research Fund).
Defining all the components of a spatially confined signaling circuit for cell movement
The team showed that the RhoA protein in the protrusions reacted to calcium influx and that RhoA activity was dependent on activation by the CaMKII protein.

Iu brought the study full circle by showing the RhoA/CaMKII calcium sensor was linked through the TEM4 protein and demonstrating that this signalling hub controlled the actin filaments pushing out the lamellipodia to the TRPV4-containing cell edges.
Further collaborations with the Tanentzapf lab at University of British Columbia revealed that Iu's results in cultured cells in vitro were also supported ex vivo in migrating mouse melanoblasts.
Iu therefore fully defined a spatially confined calcium signaling circuit that orchestrates actin cytoskeletal organization in the lamellipodia cellular domain as part of his PhD in Cell & Systems Biology.
The next steps for the Plotnikov lab are to determine how TRPV4 is activated and the clinical significance of TRPV4-dependent cell migration, particularly given the critical role of cell migration in pathological conditions such as atherosclerosis and cancer metastasis.
Dr Ernest Iu has earned the Yen fellowship at the University of Chicago to conduct his postdoctoral research under the mentorship of Professor Margaret Gardel, a world-leading expert in cell and tissue biophysics. Congratulations on this impressive work!
Professor Eyal Gruntman earns funding for cutting-edge equipment to probe the brains of flies
A new grant to Professor Eyal Gruntman from the Canadian Foundation for Innovation (CFI) will purchase a light-sheet microscope to speed up his experiments and greatly increase the area of the brain he can survey.
For his experiments at UofT Scarborough, fruit flies are suspended in a circular LED visual arena that presents visual stimuli to the flies and records their responses. The amplitude of wingbeats is measured from their shadows and that is how their turning responses are deduced.
“One test we can use relies on the reverse Phi illusion,” Gruntman explains. “If you show a bar of light moving in one direction but you rapidly flip the contrast, flies (and humans) actually perceive the bar as moving in the opposite direction. The behaviour of the fly will reflect whether it is perceiving the illusion.”
Gruntman can record the activity of the visual centres of the fly brain to assess how the behaviour of the fly is reflected in neural activity. His previous work showed that neurons responsible for computing motion change their directional preference when presented with a reverse phi stimulus.
The CFI-funded light sheet microscope (LSM) will be combined with expansion microscopy to provide unprecedented levels of detail. Expansion microscopy increases the resolution of images by physically expanding the sample and revealing details that would otherwise be below the diffraction limit of light.
Gruntman will assess how circuit function is maintained despite dynamic changes to neuronal connectivity. “We want to look downstream of motion-computing neurons,” Gruntman explains.
“In the fly, just like in mammals, there's this split where motion is computed separately for bright and dark objects, but then it's reintegrated at a particular neuropil, a bundle of nerve fibres. Neurons in the Bright and Dark pathways change their morphology throughout the day, the LSM will allow us to follow this process throughout the circuit and see how information is actually being reintegrated.”
The Gruntman lab currently has two master’s students and several undergrads currently in the lab. He is seeking additional graduate students for imaging work using the light sheet microscope made possible by CFI funding.
Congratulations, Professor Gruntman!
















